Difficult‑to‑express proteins (DEPs) include toxoids, enzymes, membrane proteins (MPs), antibodies, and some proteins with poor solubility in prokaryotic expression systems. DEPs are currently being developed as therapeutic proteins, such as drug carriers, antibodies, antimicrobial agents, and vaccines [1]. The E. coli system is widely used for DEP production because of its relative simplicity and low cost. However, many problems remain, including inherent low solubility, low yield, structural instability, and host toxicity. The water solubility and stability of DEPs mainly depend on their folding homeostasis and protein quality control (PQC); some misfolded proteins may bypass PQC mechanisms and remain in a soluble form in vivo [2].

Recently, a review published in Metabolic Engineering titled "Improving the soluble expression of difficult‑to‑express proteins in prokaryotic expression system via protein engineering and synthetic biology strategies" [3] discussed recent advances in enhancing soluble expression of DEPs in prokaryotic systems through protein engineering and synthetic biology. The article outlines key factors and mechanisms affecting proper protein folding, and summarizes advanced protein engineering techniques and tools, protein quality control systems, redesign of prokaryotic expression systems, and progress in cell‑free expression technologies for improving soluble expression of DEPs.
Approaches to Improve Soluble Expression of DEPs
✦ Protein Engineering Techniques
One of the key prerequisites for protein function is folding into the correct three‑dimensional structure. Folding is a self‑assembly process from an unfolded state to the native state, largely determined by the amino acid sequence. To date, many evolution‑based and rational design protein engineering methods have been explored to redesign the amino acid sequences of DEPs, aiming to increase protein folding stability with minimal functional interference. The charge, polarity, and interactions with surrounding molecules of amino acids, as well as pH, temperature, metal ions, cofactors, and surfactants, are all critical factors affecting the apparent stability of proteins in water or membranes [4].
Another traditional method is site‑directed mutagenesis, the most widely used engineering approach for rationally modifying DEP sequences by inserting, deleting, or substituting key residues [5]. By identifying non‑functional hydrophobic residues exposed to solvent and replacing them with hydrophilic residues, surface polar interactions can be increased, reducing their tendency to aggregate. Figure 1 shows representative techniques such as the QTY code technology, phage‑assisted directed evolution, and the PROSS method for rational protein sequence design.
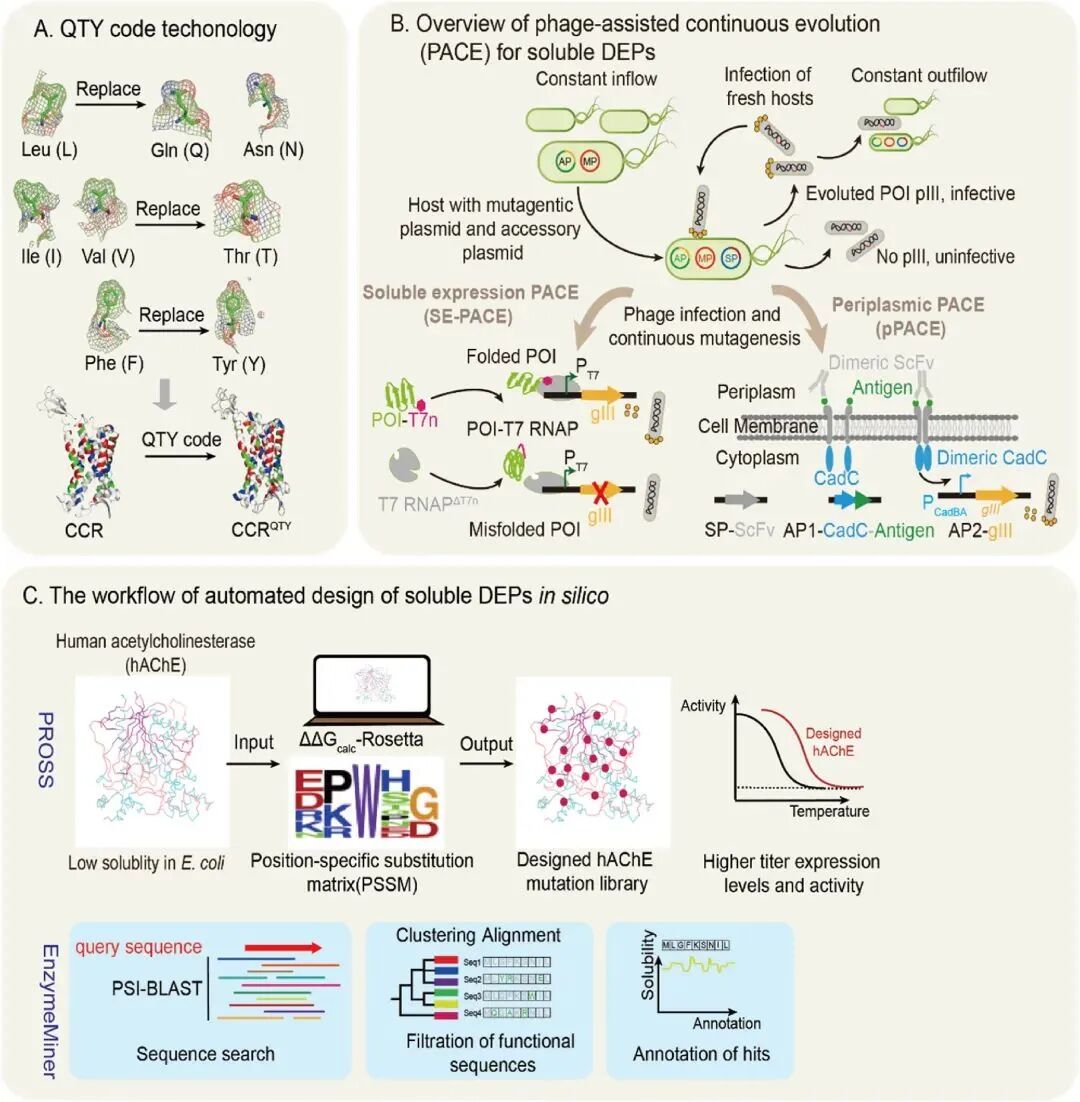
Figure 1. Representative protein engineering techniques for improving DEP solubility.
✦ Fusion with Soluble Tags
Another widely used strategy to obtain soluble proteins is fusion with soluble tags, which can simultaneously improve both solubility and yield of DEPs. Soluble tags such as glutathione S‑transferase (GST), maltose‑binding protein (MBP), small ubiquitin‑like modifier (SUMO), and Fh8 have been extensively used to enhance the soluble form of DEPs when expressed in prokaryotic hosts. Fusion partners also confer additional properties to heterologous proteins, such as protein immunodetection, quantification, and protein‑protein interactions [6].
✦ Optimization of Transcriptional and Translational Elements
Protein folding is a highly dynamic process, in which the speed of translation elongation and various co‑translational interactions play important roles. Ribosome stalling can lead to ribosome dissociation and degradation of incomplete proteins. DEP production can be regulated at multiple levels, including transcription, translation, and co‑translational folding [7].
Codon optimization techniques focus on alleviating translation elongation bottlenecks by altering low‑abundance codons, assessed using the codon adaptation index (CAI). Researchers typically eliminate rare codons by adjusting GC content and the ribosome binding site (RBS) sequence, while avoiding hairpin formation [8].
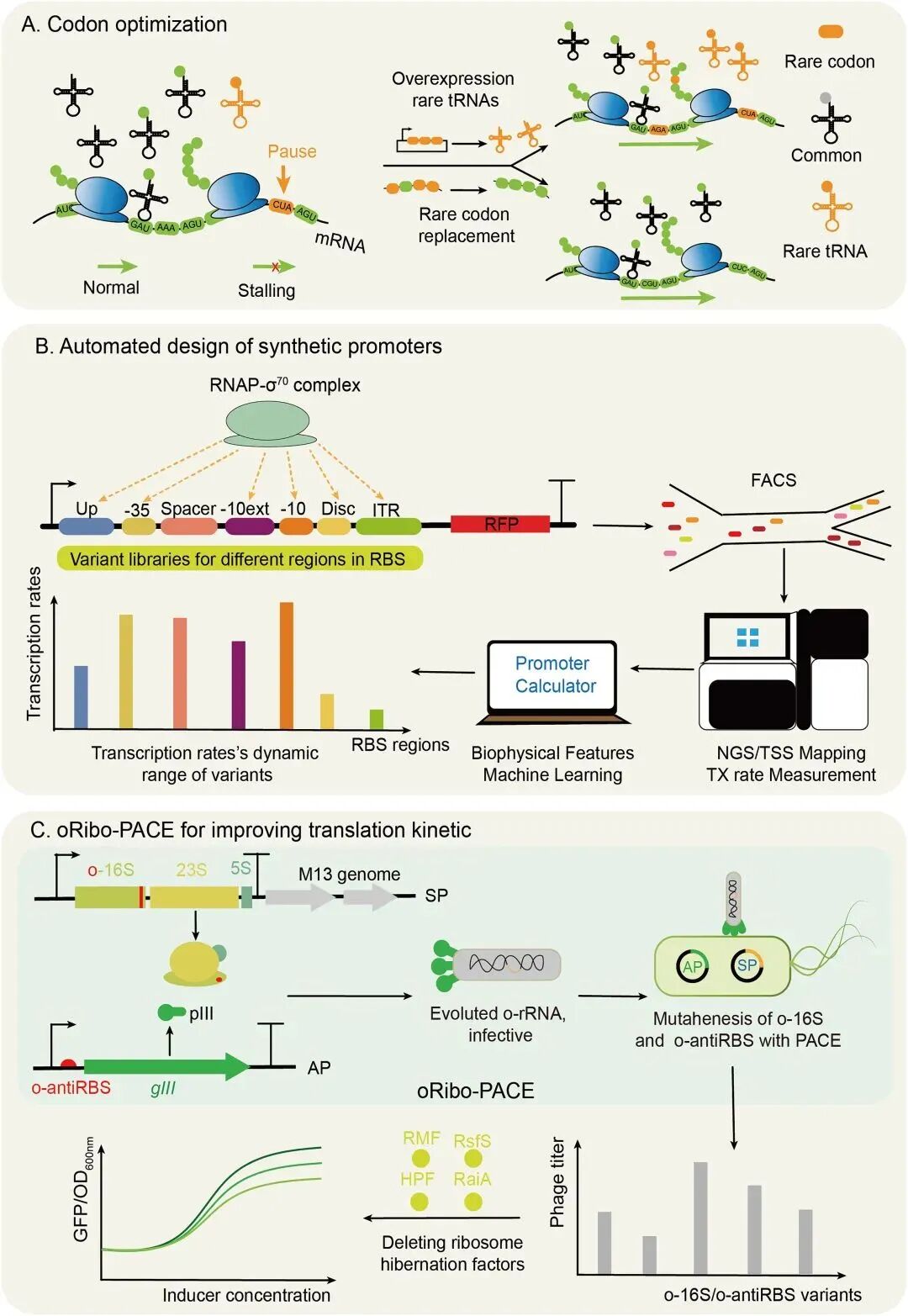
Figure 2. Optimization of transcriptional and translational elements.
✦ Co‑expression of Molecular Chaperones and Protein Quality Control (PQC) Systems
Although the development and application of soluble fusion tags are quite comprehensive, not all target proteins show consistent solubility. Therefore, co‑expression of chaperones, trigger factors, disaggregases, and disulfide isomerases is widely used as a complementary approach to improve the stability and solubility of DEPs [9].
The main function of chaperones is to recognize folding intermediates or misfolded proteins and provide a folding environment separate from the cytoplasm. The cytoplasmic chaperone systems GroEL‑GroES and DnaK‑DnaJ‑GrpE are the most thoroughly studied, and the structural basis and co‑folding mechanisms of chaperones have been extensively investigated [10].
Microorganisms also rely on chaperones and PQC systems to recognize and promote the removal of soluble misfolded proteins. Cytoprotective PQC systems have evolved the ability to selectively recognize and reduce the toxic effects of misfolded proteins to maintain protein homeostasis. Consequently, non‑functional proteins must be degraded by various bacterial PQC systems, such as ClpAP and ClpXP complexes [11].
Figure 3. Co‑expression of chaperones and PQC systems.
✦ Engineering of DEP Expression Platforms
Failure of soluble expression of DEPs, even for well‑characterized proteins, can lead to suboptimal performance of microbial cell factories, resulting in inefficient or futile research and production attempts. This problem can be overcome by expression host screening and chassis redesign.
In addition, adaptive laboratory evolution can be used to screen mutant strains for efficient DEP production. This approach does not require in‑depth knowledge of the types of mutations needed for phenotypic changes, but relies on natural selection and mutations conferring higher fitness to achieve strain optimization, such as mutants C43(DE3) and Mutant56(DE3) [12].
Adaptive laboratory evolution can induce adaptive random mutations in whole‑genome cultures to promote active bacterial growth, followed by screening to select target phenotypic mutants with better growth characteristics than the original strain.
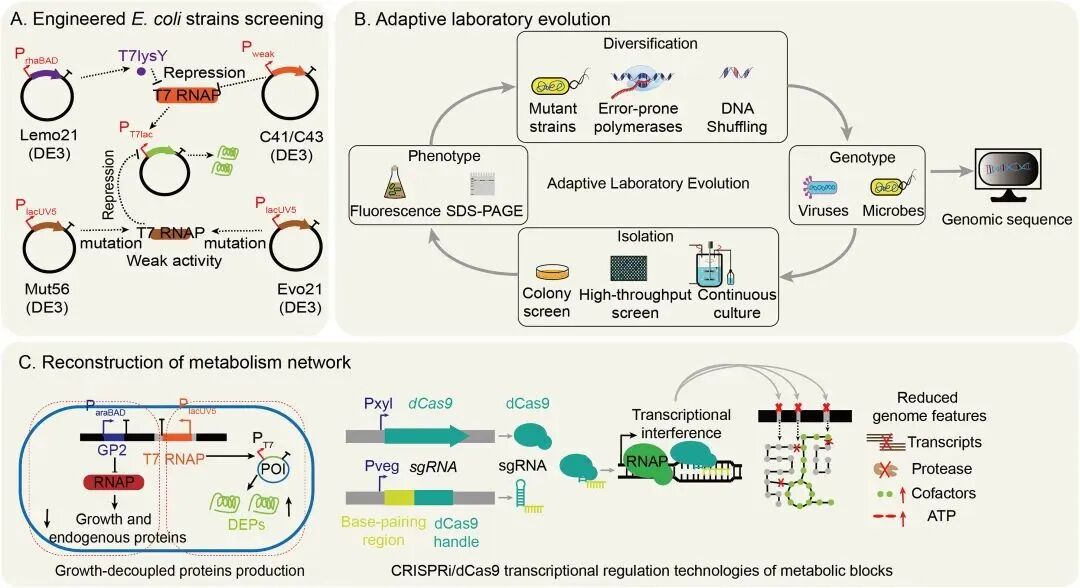
Figure 4. Overview of engineered strains for DEP expression.
✦ Cell‑Free Expression (CFE) Technology
Cell‑based recombinant protein expression can sometimes be limited by the toxicity of heterologous proteins to the host and degradation by proteases. CFE systems offer an alternative technology to circumvent the limitations of living cell expression systems. In such systems, protein synthesis can be initiated using the transcription and translation machinery in cell extracts, supplemented with substrates, DNA templates, and energy substrates required for translation [13].
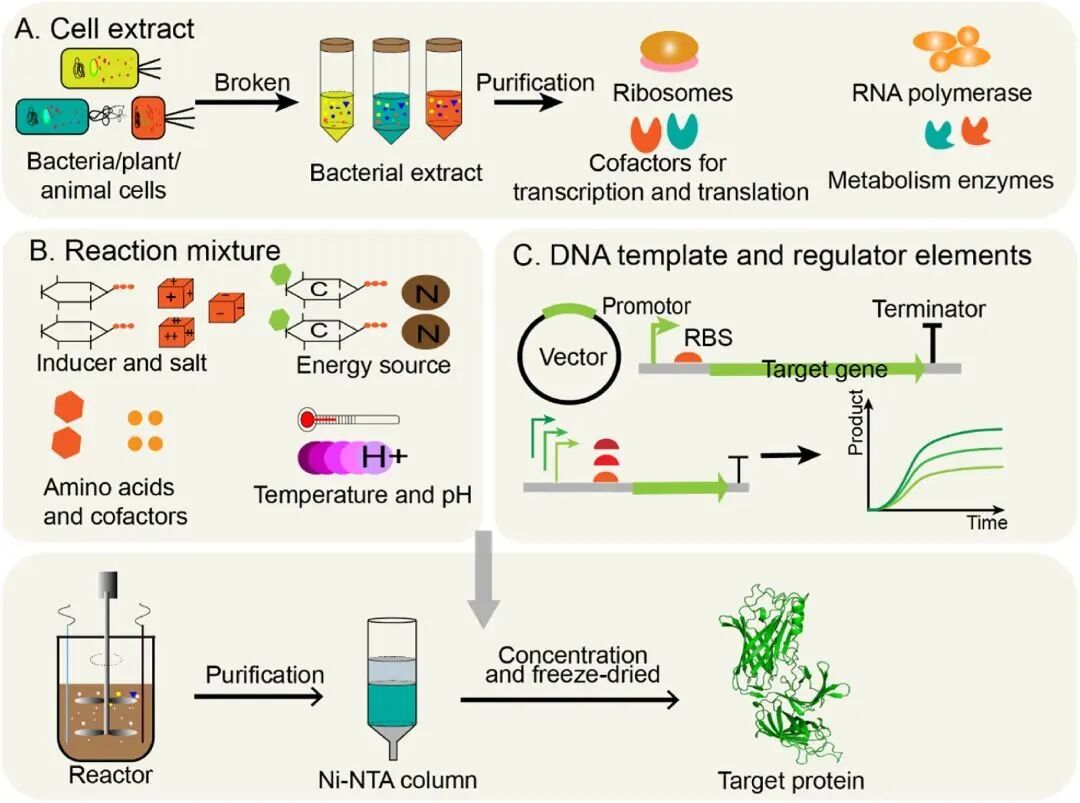
Figure 5. Workflow of cell‑free expression.
Conclusion and Outlook
In summary, prokaryotic expression systems show significant potential for scalable and cost‑effective production of high‑quality recombinant proteins. A deep understanding of protein folding mechanisms, combined with various optimization methods such as media selection, fermentation parameter adjustment, and advanced synthetic biology tools, can significantly improve the solubility and stability of DEPs. However, due to the diversity of DEPs in structure, species origin, and intrinsic properties, the combination of different approaches is often more effective than a single strategy. Moreover, with the application of machine learning, deep learning, and new technologies based on non‑natural amino acids, next‑generation cell factories and cell‑free expression systems are expected to further improve the efficiency and quality of protein production. These integrated strategies provide important new avenues for the large‑scale production of DEPs.
Looking forward, with the application of new tools, development of deep learning‑based algorithms, and introduction of non‑natural amino acids, it is expected that the de novo design and diversification capabilities for proteins will be further expanded. Given the growing attention to computational biology, deep learning‑based structure prediction, protein design, and genome‑scale metabolic models have become recent research hotspots. For example, advances in protein structure prediction have made it possible to improve protein solubility by accurately identifying and reducing surface hotspot regions without affecting core interactions or risking structural collapse. On the other hand, the emergence of novel metabolic network models is expected to enable next‑generation cell factories with high expression capacity.
Canton Biologics' DTEasy Platform
Canton Biologics has developed the DTEasy toolbox for difficult‑to‑express protein drugs. This platform integrates and optimizes each key stage of the DEP expression process, successfully increasing protein yield and helping protein drug molecules move smoothly toward industrialization. Specific details of the Canton Biologics DEP platform will be covered in a subsequent introduction.
References
[1] Grund, M.E., Soo, J.C., Cote, C.K., Berisio, R., Lukomski, S., 2021. Thinking outside the bug: targeting outer membrane proteins for burkholderia vaccines. Cells 10 (3), 495.
[2] Nissley, D.A., Jiang, Y., Trovato, F., Sitarik, I., Narayan, K.B., To, P., Xia, Y., Fried, S.D., O’Brien, E.P., 2022. Universal protein misfolding intermediates can bypass the proteostasis network and remain soluble and less functional. Nat. Commun. 13 (1), 3081.
[3] JinPing C ,JinSong G ,Chang S , et al.Improving the soluble expression of difficult-to-express proteins in prokaryotic expression system via protein engineering and synthetic biology strategies.[J].Metabolic engineering,2023,7899-114.
[4] Kaur, J., Kumar, A., Kaur, J., 2018. Strategies for optimization of heterologous protein expression in E. coli: roadblocks and reinforcements. Int. J. Biol. Macromol. 106, 803–822.
[5] Ali, S.A., Hassan, M.I., Islam, A., Ahmad, F., 2014. A review of methods available to estimate solvent-accessible surface areas of soluble proteins in the folded and unfolded states [Review]. Curr. Protein Pept. Sci. 15 (5), 456–476.
[6] Fox, J.D., Routzahn, K.M., Bucher, M.H., Waugh, D.S., 2003. Maltodextrin-binding proteins from diverse bacteria and archaea are potent solubility enhancers. FEBS Lett. 537 (1–3), 53–57.
[7] Presnyak, V., Alhusaini, N., Chen, Y.H., Martin, S., Morris, N., Kline, N., Olson, S., Weinberg, D., Baker, K.E., Graveley, B.R., Coller, J., 2015. Codon optimality is a major determinant of mRNA stability. Cell 160 (6), 1111–1124.
[8] Bradley, R.W., Buck, M., Wang, B., 2016. Tools and principles for microbial gene circuit engineering. J. Mol. Biol. 428 (5), 862–888.
[9] Oguchi, Y., Kummer, E., Seyffer, F., Berynskyy, M., Anstett, B., Zahn, R., Wade, R.C., Mogk, A., Bukau, B., 2012. A tightly regulated molecular toggle controls AAA+ disaggregase. Nat. Struct. Mol. Biol. 19 (12), 1338–1346.
[10] Hayer-Hartl, M., Bracher, A., Hartl, F.U., 2016. The GroEL–GroES chaperonin machine: a nano-cage for protein folding. Trends Biochem. Sci. 41 (1), 62–76.
[11] Fei, X., Bell, T.A., Barkow, S.R., Baker, T.A., Sauer, R.T., 2020. Structural basis of ClpXP recognition and unfolding of ssrA-tagged substrates. Elife 9. Article e61496.
[12] Gheibi Hayat, S.M., Farahani, N., Golichenari, B., Sahebkar, A., 2018. Recombinant protein expression in E. coli: what we need to know. Curr. Pharmaceut. Des. 24, 718–725.
[13] Sandberg, T.E., Salazar, M.J., Weng, L.L., Palsson, B.O., Feist, A.M., 2019. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab. Eng. 56, 1–16。

